Category Archive 'Genetics'
22 Jan 2018

The Conversation seems to have an explanation for how we got millennials.
The Y chromosome may be a symbol of masculinity, but it is becoming increasingly clear that it is anything but strong and enduring. Although it carries the “master switch†gene, SRY, that determines whether an embryo will develop as male (XY) or female (XX), it contains very few other genes and is the only chromosome not necessary for life. Women, after all, manage just fine without one.
What’s more, the Y chromosome has degenerated rapidly, leaving females with two perfectly normal X chromosomes, but males with an X and a shrivelled Y. If the same rate of degeneration continues, the Y chromosome has just 4.6m years left before it disappears completely. This may sound like a long time, but it isn’t when you consider that life has existed on Earth for 3.5 billion years.
RTWT
25 Jul 2017


Homo erectus imagined by the Smithsonian.
Before It’s News:
The evolutionary history of a salivary protein may point to interbreeding between humans and an enigmatic ancient relative
In saliva, scientists have found hints that a “ghost†species of archaic humans may have contributed genetic material to ancestors of people living in Sub-Saharan Africa today.
The research adds to a growing body of evidence suggesting that sexual rendezvous between different archaic human species may not have been unusual.
Past studies have concluded that the forebears of modern humans in Asia and Europe interbred with other early hominin species, including Neanderthals and Denisovans. The new research is among more recent genetic analyses indicating that ancient Africans also had trysts with other early hominins.
“It seems that interbreeding between different early hominin species is not the exception — it’s the norm,†says Omer Gokcumen, PhD, an assistant professor of biological sciences in the University at Buffalo College of Arts and Sciences.
“Our research traced the evolution of an important mucin protein called MUC7 that is found in saliva,†he says. “When we looked at the history of the gene that codes for the protein, we see the signature of archaic admixture in modern day Sub-Saharan African populations.â€
The research was published on July 21 in the journal Molecular Biology and Evolution. The study was led by Gokcumen and Stefan Ruhl, DDS, PhD, a professor of oral biology in UB’s School of Dental Medicine.
The scientists came upon their findings while researching the purpose and origins of the MUC7 protein, which helps give spit its slimy consistency and binds to microbes, potentially helping to rid the body of disease-causing bacteria.
As part of this investigation, the team examined the MUC7 gene in more than 2,500 modern human genomes. The analysis yielded a surprise: A group of genomes from Sub-Saharan Africa had a version of the gene that was wildly different from versions found in other modern humans.
The Sub-Saharan variant was so distinctive that Neanderthal and Denisovan MUC7 genes matched more closely with those of other modern humans than the Sub-Saharan outlier did.
“Based on our analysis, the most plausible explanation for this extreme variation is archaic introgression — the introduction of genetic material from a ‘ghost’ species of ancient hominins,†Gokcumen says. “This unknown human relative could be a species that has been discovered, such as a subspecies of Homo erectus, or an undiscovered hominin. We call it a ‘ghost’ species because we don’t have the fossils.â€
Given the rate that genes mutate during the course of evolution, the team calculated that the ancestors of people who carry the Sub-Saharan MUC7 variant interbred with another ancient human species as recently as 150,000 years ago, after the two species’ evolutionary path diverged from each other some 1.5 to 2 million years ago.
RTWT
03 Jun 2016


The Atlantic has news about the (dual) origin of man’s best friend.
On the eastern edge of Ireland lies Newgrange, a 4,800-year-old monument that predates Stonehenge and the pyramids of Giza. Beneath its large circular mound and within its underground chambers lie many fragments of animal bones. And among those fragments, Dan Bradley from Trinity College Dublin found the petrous bone of a dog.
Press your finger behind your ear. That’s the petrous. It’s a bulbous knob of very dense bone that’s exceptionally good at preserving DNA. If you try to pull DNA out of a fossil, most of it will come from contaminating microbes and just a few percent will come from the bone’s actual owner. But if you’ve got a petrous bone, that proportion can be as high as 80 percent. And indeed, Bradley found DNA galore within the bone, enough to sequence the full genome of the long-dead dog.
Larson and his colleague Laurent Frantz then compared the Newgrange sequences with those of almost 700 modern dogs, and built a family tree that revealed the relationships between these individuals. To their surprise, that tree had an obvious fork in its trunk—a deep divide between two doggie dynasties. One includes all the dogs from eastern Eurasia, such as Shar Peis and Tibetan mastiffs. The other includes all the western Eurasian breeds, and the Newgrange dog.
The genomes of the dogs from the western branch suggest that they went through a population bottleneck—a dramatic dwindling of numbers. Larson interprets this as evidence of a long migration. He thinks that the two dog lineages began as a single population in the east, before one branch broke off and headed west. This supports the idea that dogs were domesticated somewhere in China.
But there’s a critical twist.
The team calculated that the two dog dynasties split from each other between 6,400 and 14,000 years ago. But the oldest dog fossils in both western and eastern Eurasia are older than that. Which means that when those eastern dogs migrated west into Europe, there were already dogs there.
Here’s the full story, as he sees it. Many thousands of years ago, somewhere in western Eurasia, humans domesticated grey wolves. The same thing happened independently, far away in the east. So, at this time, there were two distinct and geographically separated groups of dogs. Let’s call them Ancient Western and Ancient Eastern. Around the Bronze Age, some of the Ancient Eastern dogs migrated westward alongside their human partners, separating from their homebound peers and creating the deep split in Larson’s tree. Along their travels, these migrants encountered the indigenous Ancient Western dogs, mated with them (doggy style, presumably), and effectively replaced them.
Today’s eastern dogs are the descendants of the Ancient Eastern ones. But today’s western dogs (and the Newgrange one) trace most of their ancestry to the Ancient Eastern migrants. Less than 10 percent comes from the Ancient Western dogs, which have since gone extinct.
Read the whole thing.

Newgrange, the white facade is modern.
13 Dec 2015


Benjamin Jowett 1817-1893
My name is Benjamin Jowett.
I am Master of Balliol College.
What there is to know, I know it,
And what I don’t know isn’t knowledge.
Nobody would write a poem like that about Yale’s current president Peter Salovey.
————————-
A recent paper in the journal Intelligence is titled:
Were the Victorians cleverer than us? The decline in general intelligence estimated from a meta-analysis of the slowing of simple reaction time
Highlights
• Simple reaction time has slowed since 1889.
• Psychometric meta-analysis reveals a decline in g of − 1.16 points per decade.
• The decline between 1889 and 2004 is − 13.35 points.
• The decline between 1889 and 2004 is − 12.45 points.
• This is the first direct measurement of a probable dysgenic trend in IQ.
Abstract:
The Victorian era was marked by an explosion of innovation and genius, per capita rates of which appear to have declined subsequently. The presence of dysgenic fertility for IQ amongst Western nations, starting in the 19th century, suggests that these trends might be related to declining IQ. This is because high-IQ people are more productive and more creative. We tested the hypothesis that the Victorians were cleverer than modern populations, using high-quality instruments, namely measures of simple visual reaction time in a meta-analytic study. Simple reaction time measures correlate substantially with measures of general intelligence (g) and are considered elementary measures of cognition. In this study we used the data on the secular slowing of simple reaction time described in a meta-analysis of 14 age-matched studies from Western countries conducted between 1889 and 2004 to estimate the decline in g that may have resulted from the presence of dysgenic fertility. Using psychometric meta-analysis we computed the true correlation between simple reaction time and g, yielding a decline of − 1.16 IQ points per decade or − 13.35 IQ points since Victorian times. These findings strongly indicate that with respect to g the Victorians were substantially cleverer than modern Western populations.
24 Jul 2015

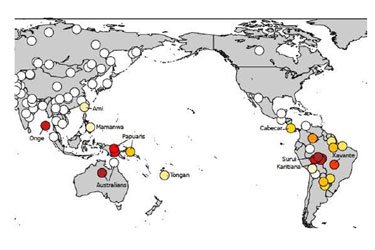
The Register describes the findings of genetic research reported in a paper in Nature, titled “Genetic evidence for two founding populations of the Americas.”
A long-vanished race of humans, whose descendants now survive only among certain indigenous peoples in Australasia and in the Amazon jungles, may have been the true, original Native Americans, according to new genetics research.
The clue to the existence of this mysterious “Population Y” has been found by boffins probing the genes of various native-American peoples. Amazingly, the researchers found that a couple of tribes in the Amazon jungle actually had quite a lot in common genetically with others in Australia, New Guinea and the Andaman Islands.
It’s well known, of course, that humanity spread from its original cradle in Africa out across Asia and thus south to Australasia and separately north – via the Bering Strait land bridge, then in existence – to North America.
The first emigrants across the Bering were the First Americans. What’s not totally clear is exactly when they arrived, how many of them there were, what other groups arrived at what point etc. Given that they were arriving via a travel route just south of the ice sheets which then covered a lot of the northern globe, it’s generally thought that they didn’t come in a continuous stream, but more probably in waves when conditions were favourable.
One widely-held theory has it that the First Americans arrived in one wave of fairly genetically similar people, and this theory fitted pretty well with the observed genetics of native-American people both North and South. But the Amazon-Australasia linkage has pretty much upset that applecart.
“We spent a really long time trying to make this result go away and it just got stronger,” says Professor David Reich of the Harvard Medical School.
Read the whole thing.
Unmentioned besides is a possible third group, the Solutrean Hypothesis, a theory based on archaeological evidence which proposes also some Trans-Atlantic settlement of the Americas.
08 Jul 2015


Adam Piore, in Nautilus, tells the fascinating story of Decode, the company founded in 1996 to collect and study Icelandic DNA, using the genetics of that island nations’s small and closely-related population to find genetic links to common diseases.
In the ninth century there was a Norwegian Viking named Kveldulf, so big and strong that no man could defeat him. He sailed the seas in a long-ship and raided and plundered towns and homesteads of distant lands for many years. He settled down to farm, a very wealthy man.
Kveldulf had two sons who grew up to become mighty warriors. One joined the service of King Harald Tangle Hair. But in time the King grew fearful of the son’s growing power and had him murdered. Kveldulf vowed revenge. With his surviving son and allies, Kveldulf caught up with the killers, and wielding a double-bladed ax, slew 50 men. He sent the paltriest survivors back to the king to recount his deed and fled toward the newly settled realm of Iceland. Kveldulf died on the journey. But his remaining son Skallagrim landed on Iceland’s west coast, prospered, and had children.
Skallagrim’s children had children. Those children had children. And the blood and genes of Kveldulf the Viking and Skallagrim his son were passed down the ages. Then, in 1949, in the capital of Reykjavik, a descendent named Kari Stefansson was born.
Like Kveldulf, Stefansson would grow to be a giant, 6’5â€, with piercing eyes and a beard. As a young man, he set out for the distant lands of the universities of Chicago and Harvard in search of intellectual bounty. But at the dawn of modern genetics in the 1990s, Stefansson, a neurologist, was lured back to his homeland by an unlikely enticement—the very genes that he and his 300,000-plus countrymen had inherited from Kveldulf and the tiny band of settlers who gave birth to Iceland.
Stefansson had a bold vision. He would create a library of DNA from every single living descendent of his nation’s early inhabitants. This library, coupled with Iceland’s rich trove of genealogical data and meticulous medical records, would constitute an unparalleled resource that could reveal the causes—and point to cures—for human diseases.
Read the whole thing.
22 Jan 2013


90% of living Jews are European (Askenazi) Jews. There are two theories of the origin of European Jewry: the Rhineland Hypothesis contends that European Jews fled Palestine after the Roman destruction of the Temple in 70 A.D. or after the Muslim conquest of Palestine in 637 A.D., migrated into Europe via Italy and Spain, then settled along the Rhine, before being driven eastward by persecution. The use of Yiddish, a High German dialect, by Ashkenazi Jews provides substantive support for their origin somewhere in Germany.
The alternative Khazar Hypothesis, popularized by Arthur Koestler, argues that nearly all European Jews really descend from the Khazars, a Turkish-speaking people, who converted to Judaism en masse in the 8th century.
New genetic evidence produced by a study by Geneticist Eran Elhaik of the Johns Hopkins School of Public Health in Baltimore, published in the British journal Genome Biology and Evolution, on The Missing Link of Jewish European Ancestry: Contrasting the Rhineland and the Khazarian Hypotheses provides strong support for the Khazar Hypothesis.
Abstract excerpt
Thus far… the Khazars’ contribution has been estimated only empirically, as the absence of genome-wide data from Caucasus populations precluded testing the Khazarian hypothesis. Recent sequencing of modern Caucasus populations prompted us to revisit the Khazarian hypothesis and compare it with the Rhineland hypothesis. We applied a wide range of population genetic analyses to compare these two hypotheses. Our findings support the Khazarian hypothesis and portray the European Jewish genome as a mosaic of Near Eastern-Caucasus, European, and Semitic ancestries, thereby consolidating previous contradictory reports of Jewish ancestry. We further describe a major difference among Caucasus populations explained by the early presence of Judeans in the Southern and Central Caucasus.
——————————–
In the same issue, Danielle Venton summarized the conclusion of “the first scientific paper to prove the Khazarian Hypothesis and reject the Rhineland Hypothesis.”
When Behar et al. published “The genome-wide structure of the Jewish people†in 2010, Elhaik decided to investigate the question that had intrigued him for so long. Using data published by Behar, he calculated seven measures of ancestry, relatedness, and geographical origin. Though he used some of the same statistical tests as prior studies, he chose different comparisons.
“Results in the current literature are tangled,†Elhaik says. “Everyone is basically following the same assumption: Ashkenazi Jews are a population isolate, so they are all similar to one another, and this is completely incorrect.â€
Previous studies had, for example, combined the question of similarity among and between Jewish populations and the question of ancestry and relatedness to non-Jewish populations. Elhaik viewed these questions separately. Jewish communities are less homogeneous than is popularly thought, he says, with Jewish communities along the former Khazarian border showing the most heterogeneity.
His second question centered on ancestry: When comparing Jewish communities to their non-Jewish neighbors, Caucasus or Levant (Middle Eastern) populations—which is the closest to Jews? “All Eurasian Jewish communities are closer to Caucasus populations,†he writes, with Central European Jews closer to Italian non-Jews as the exception. Not one of the eight evaluated Jewish populations were closer to Levant populations.
29 Feb 2012


Science Daily reports that the count of blood group proteins has been increased for the first time in more than a decade by two more, raising the count from 30 to 32.
You probably know your blood type: A, B, AB or O. You may even know if you’re Rhesus positive or negative. But how about the Langereis blood type? Or the Junior blood type? Positive or negative? Most people have never even heard of these.
Yet this knowledge could be “a matter of life and death,” says University of Vermont biologist Bryan Ballif.
While blood transfusion problems due to Langereis and Junior blood types are rare worldwide, several ethnic populations are at risk, Ballif notes. “More than 50,000 Japanese are thought to be Junior negative and may encounter blood transfusion problems or mother-fetus incompatibility,” he writes.
But the molecular basis of these two blood types has remained a mystery — until now.
In the February issue of Nature Genetics, Ballif and his colleagues report on their discovery of two proteins on red blood cells responsible for these lesser-known blood types.
Ballif identified the two molecules as specialized transport proteins named ABCB6 and ABCG2.
“Only 30 proteins have previously been identified as responsible for a basic blood type,” Ballif notes, “but the count now reaches 32.”
The last new blood group proteins to be discovered were nearly a decade ago, Ballif says, “so it’s pretty remarkable to have two identified this year.”
16 Nov 2009

Are you a genetically a dandelion or an orchid? Both have their place in the evolutionary scheme of things according to a recent article in the Atlantic by David Dobbs.
Most of us have genes that make us as hardy as dandelions: able to take root and survive almost anywhere. A few of us, however, are more like the orchid: fragile and fickle, but capable of blooming spectacularly if given greenhouse care. So holds a provocative new theory of genetics, which asserts that the very genes that give us the most trouble as a species, causing behaviors that are self-destructive and antisocial, also underlie humankind’s phenomenal adaptability and evolutionary success. With a bad environment and poor parenting, orchid children can end up depressed, drug-addicted, or in jail—but with the right environment and good parenting, they can grow up to be society’s most creative, successful, and happy people.
Hat tip to Bird Dog.
17 Jul 2008


Lichtensteinhöhle skeletons
British newspapers report that living residents of Nienstedt, a village in the foothills of the Harz Mountains in Lower Saxony, have been found by DNA analysis to be relatives of 3000-year-old Bronze Age inhabitants of the same area interred in the nearby Lichtensteinhöhle cave.
—————————————————————
London Times:
The good news for two villagers in the Söse valley of Germany yesterday was that they have discovered their (127th times)-great grandparents.
The bad news is that their long-lost ancestors may have grilled and eaten other members of their clan.
Every family has its skeletons in the cave, though, so Manfred Hucht-hausen, 58, a teacher, and 48-year-old surveyor Uwe Lange remained in celebratory mood. Thanks to DNA testing of remarkably well-preserved Bronze Age bones, they can claim to have the longest proven family tree in the world. “I can trace my family back by name to 1550,†Mr Lange said. “Now I can go back 120 generations.â€
Mr Lange comes from the village of Nienstedt, in Lower Saxony, in the foothills of the Harz mountain range. “We used to play in these caves as kids. If I’d known that there were 3,000-year-old relatives buried there I wouldn’t have set foot in the place.â€
The cave, the Lichtensteinhöhle, is made up of five interlocked natural chambers. It stayed hidden from view until 1980 and was not researched properly until 1993. The archaeologist Stefan Flindt found 40 skeletons along with what appeared to be cult objects. …
Analysis showed that all the bones were from the same family and the scientists speculated that it was a living area and a ceremonial burial place.
About 300 locals agreed to giving saliva swabs. Two of the cave family had a very rare genetic pattern – and a match was found.
—————————————————————
Telegraph:
The bones of 40 people were shielded from the elements by calcium deposits that formed a protective skin around the skeletons.
All the remains turned out to be from the same family group who had a distinctive – and rare – DNA pattern.
When people in the local area were tested with saliva swabs, two nearby residents turned out to have the same distinctive genetic characteristic.
Manfred Huchthausen, a 58-year-old teacher, and Uwe Lange, a 48-year-old surveyer, now believe they are even more local than either of them thought.
—————————————————————
Inma Pazos at iGENEA Forum provides more specific information.
(translated & abridged)
DNA analysis really found that 15 of 22 skeletons were relatives, constituting several generations of a family clan. In 2007, about 300 DNA samples of today’s indigenous population in Osterode-am-Harz were collected and tested for possible affinity. Susann Hummel, a leading anthropologist, has identified eleven living persons as descendants of the cave burials.
Ten lines of mtDNA haplogroup H, four of haplogroup U, two of the haplogroup J and three of the haplogroup T were identified. A further breakdown in the sub-groups succeeded in identifying U5b, T2 and J1b1. In another case, membership in sub-group U2 was considered very likely.
—————————————————————
mtDNA haplogroups
17 Sep 2007
That old goat chasing the young girl is doing mankind a favor, researchers from Stanford and the University of California at Santa Barbara tell us.
ScienceDaily
/div>

Feeds
|














